By the time authors – who have acknowledged third-party financial support in their research papers submitted to a journal using the Pensoft-developed publishing platform: ARPHA – open their inboxes to the congratulatory message that their work has just been published and made available to the wide world, a similar notification will have also reached their research funder.
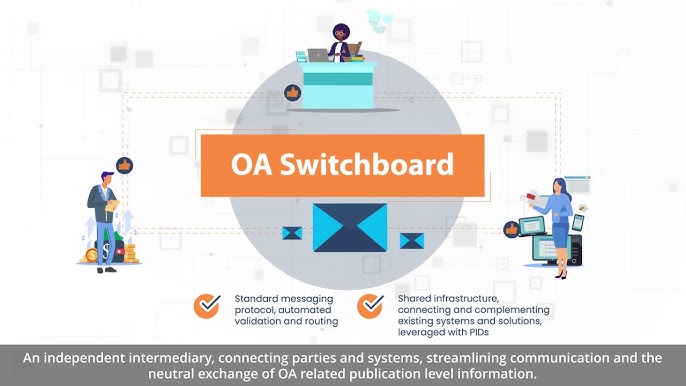
This automated workflow is already in effect at all journals (co-)published by Pensoft and those published under their own imprint on the ARPHA Platform, as a result of the new partnership with the OA Switchboard: a community-driven initiative with the mission to serve as a central information exchange hub between stakeholders about open access publications, while making things simpler for everyone involved.
All the submitting author needs to do to ensure that their research funder receives a notification about the publication is to select the supporting agency or the scientific project (e.g. a project supported by Horizon Europe) in the manuscript submission form, using a handy drop-down menu. In either case, the message will be sent to the funding body as soon as the paper is published in the respective journal.
“At Pensoft, we are delighted to announce our integration with the OA Switchboard, as this workflow is yet another excellent practice in scholarly publishing that supports transparency in research. Needless to say, funding and financing are cornerstones in scientific work and scholarship, so it is equally important to ensure funding bodies are provided with full, prompt and convenient reports about their own input.”
comments Prof Lyubomir Penev, CEO and founder of Pensoft and ARPHA.
“Research funders are one of the three key stakeholder groups in OA Switchboard and are represented in our founding partners. They seek support in demonstrating the extent and impact of their research funding and delivering on their commitment to OA. It is great to see Pensoft has started their integration with OA Switchboard with a focus on this specific group, fulfilling an important need,”
adds Yvonne Campfens, Executive Director of the OA Switchboard.
***
About the OA Switchboard:
A global not-for-profit and independent intermediary established in 2020, the OA Switchboard provides a central hub for research funders, institutions and publishers to exchange OA-related publication-level information. Connecting parties and systems, and streamlining communication and the neutral exchange of metadata, the OA Switchboard provides direct, indirect and community benefits: simplicity and transparency, collaboration and interoperability, and efficiency and cost-effectiveness.
About Pensoft:
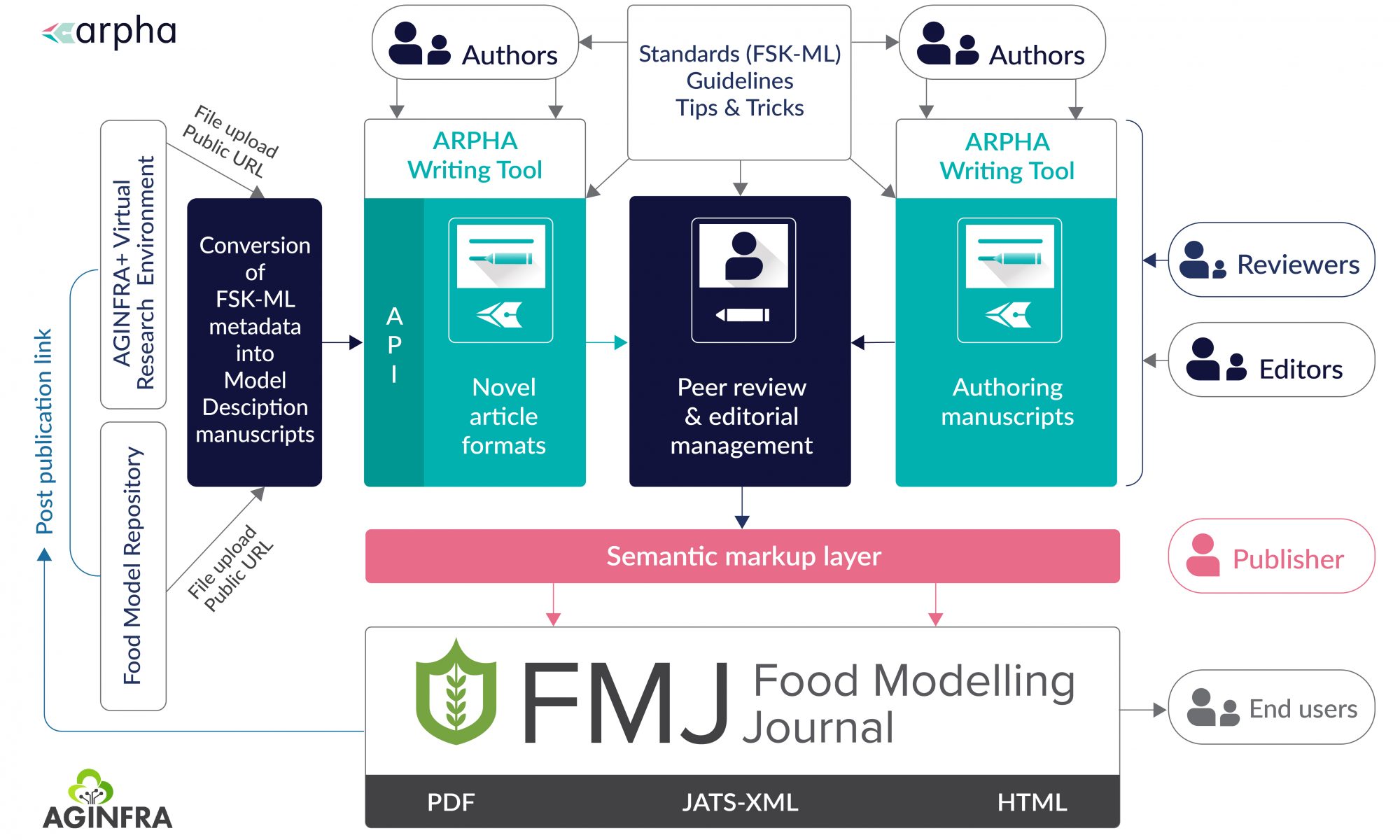
Pensoft is an independent academic publishing company, well known worldwide for its novel cutting-edge publishing tools, workflows and methods for text and data publishing of journals, books and conference materials.
All journals (co-)published by Pensoft are hosted on Pensoft’s full-featured ARPHA Publishing Platform and published in a way that ensures their content is as FAIR as possible, meaning that it is effortlessly readable, discoverable, harvestable, citable and reusable by both humans and machines.
***
Follow Pensoft on Twitter, Facebook and Linkedin.
Follow OA Switchboard on Twitter and Linkedin.
***
Image credit: OA Switchboard.