Acta Biologica Sibirica is an open-access, peer-reviewed journal on the biodiversity of Siberia and the adjacent lands by Altai State University. Since 2015, it has been publishing original research in the field of experimental and field biology.
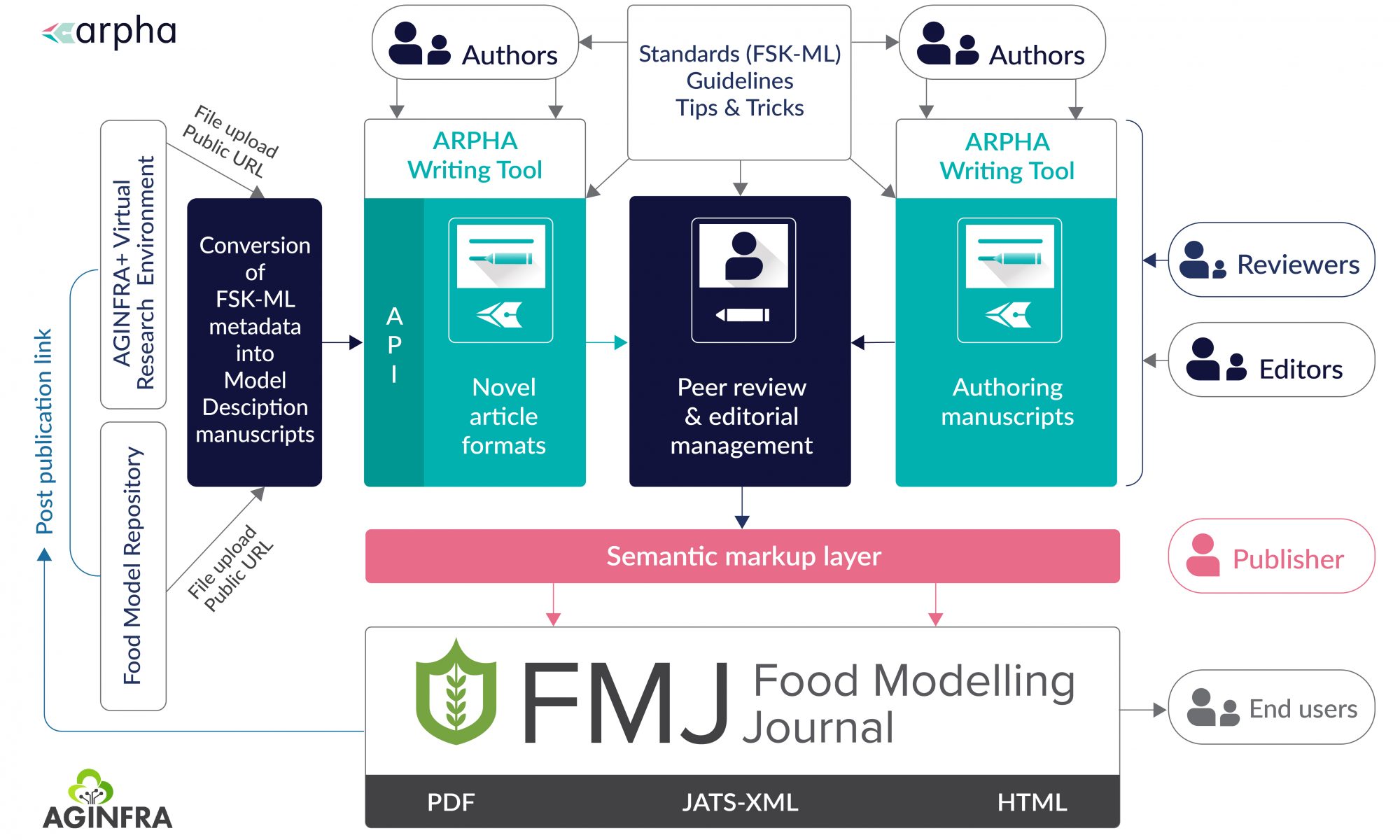
Starting from 2020, Acta Biologica Sibirica moves to the full-featured technologically advanced platform ARPHA and will be published in collaboration with the scholarly publisher and technology provider Pensoft.
Pensoft’s original publishing system ARPHA allows Acta Biologica Sibirica to publish original research papers, reviews, short communications, letters and discussion papers, book reviews and memorial articles. The scholarly platform was designed to facilitate authors in the manuscript writing, submission and review process as end-to-end experience, including publication of the data and multimedia content in the form, suitable for both enhanced high-tech human and machine discoverability of the scholarly outputs.
Acta Biologica Sibirica accepts for publication papers in taxonomy, phylogeny, biogeography, faunistics, floristics, biological systematics, nature conservation and protected areas. In the fields of faunistics and floristics there are several types of articles, available for submission: floral and faunal lists on any region of the world, faunal and floral discoveries (e.g. species newly recorded in a particular region, additions to previously published inventories), papers on methodology of faunal and floral studies.
«Our basic task is to turn our journal into a high-quality world-class publication. Without the help of modern publishers, this is almost impossible. The choice of the publisher was perfectly logical – the reputation of Pensoft Publishers and its founder, the famous Bulgarian zoologist Lubomir Penev, is impeccable. To stand in one cohort with powerful publications with a long history is an honor for us. High standards of editing and reviewing manuscripts, the absolute level of originality and scientific novelty – these are the criteria on which we will rely»,
comments the editor-in-chief of the journal, Professor of Altai University Roman Yakovlev.
«At Pensoft, we are delighted to initiate this wonderful partnership with yet another renowned research institution in Russia, namely Altai State University. With our long-year experience in zoological and biodiversity research publishing and dissemination, I am certain that the journal has found a fitting place in the family of Pensoft and ARPHA»,
comments Prof. Lyubomir Penev, CEO and founder of Pensoft and ARPHA.
The first papers of 2020 are already available online on the new website of Acta Biologica Sibirica.
Within the pioneering papers published in the renewed journal, there is a research article about the first result of DNA-studies on the Central Asiatic owlet moths in the genus Euchalcia. The studied specimens were collected in Kyrgyzstan and Kazakhstan during the expeditions of the Russian Entomological Society in 2017-2019. When comparing a specific mitochondrial gene (cytochrome C oxidase subunit I or COI) between various species, the scientists revealed that the difference amongst European Euchalcia species is smaller than the one amongst high-mountainous Central Asiatic species.
Another study records the first occurrence of the moorland clouded yellow in Altai Region. The butterfly was found to share a mitochondrial barcode with some specimens from mountain populations from the Alps and the Czech Republic.

Credit: Nazar A. Shapoval
License: CC-BY 4.0
***
Follow Acta Biologica Sibirica on Twitter and Facebook.
***
Additional information
About Altai State University:
Altai State University is one of the leading Russian classical higher education institutions established in 1973. It is a major educational, research and cultural center located in the Asian part of the country, integrated into the international academic community, training the intellectual elite and conducting high-impact research.
The unique geographical position of Altai region, located in the center of Asia predestinates the University’s mission – to appear as an international research and educational center that integrates, develops and spreads the modern Western, Russian and Asian knowledge in education, science and culture within the Asian region.
About ARPHA:
ARPHA is the first end-to-end, narrative- and data-integrated publishing solution that supports the full life cycle of a manuscript, from authoring to reviewing, publishing and dissemination. ARPHA provides accomplished and streamlined production workflows that can be customized according to the journal’s needs. The platform enables a variety of publishing models through a number of options for branding, production and revenue models to choose from.
About Pensoft:
Pensoft is an independent academic publishing company, well-known worldwide for its innovations in the field of semantic publishing, as well as for its cutting-edge publishing tools and workflows. In 2013, Pensoft launched the first ever end to end XML-based authoring, reviewing and publishing workflow, as demonstrated by the Pensoft Writing Tool (PWT) and the Biodiversity Data Journal (BDJ), now upgraded to the ARPHA Publishing Platform. Flagship titles include: Research Ideas and Outcomes (RIO), One Ecosystem, ZooKeys, Biodiversity Data Journal, PhytoKeys, MycoKeys and many more.
Contacts:
Prof. Lyubomir Penev, founder and CEO at ARPHA and Pensoft
Email: penev@pensoft.net
Prof. Alex Matsyura, Editor-in-Chief of Acta Biologica Sibirica
Email: amatsyura@gmail.com
Prof. Roman Yakovlev, Editor-in-Chief of Acta Biologica Sibirica
Email: yakovlev_asu@mail.ru